Original Article Details:
Title: PAM-IndependentCRISPR-Cas12aSystemforSpecificAssaysof Single Nucleotide Variants
Authors: JinlongAi, JinhaiDeng, JingjingHu, XingxiangPu, *TongyanYuan, YulingTeng, HanLi, BojieChen, JinlianDu, LingJiang, *XiaoyanChen, ErhuXiong, *and RonghuaYang.
Journal: JACS Au
DOI: https://doi.org/10.1021/jacsau.5c00011
This study introduces TRACER, a PAM-independent CRISPR-Cas12a detection method that overcomes a major limitation of CRISPR diagnostics. By converting amplified double-stranded DNA into single-stranded DNA, TRACER allows Cas12a to detect targets without requiring PAM sequences. The system achieves ultra-sensitive detection down to attomolar levels and can identify rare mutations present at just 0.1% abundance, with strong validation in clinical cancer samples.
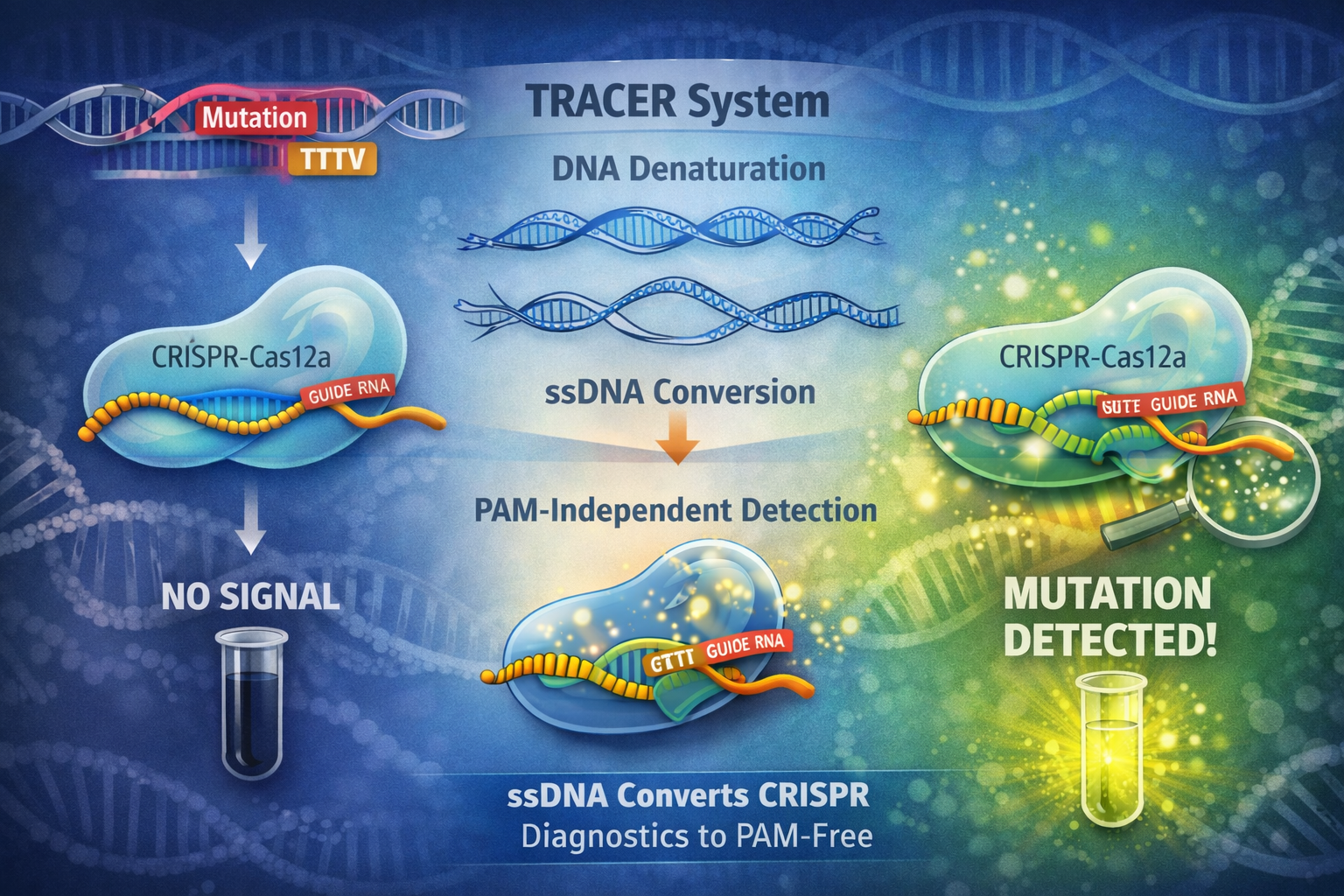
The Problem No One Notices — Until It Breaks the Experiment
In modern molecular diagnostics, CRISPR-Cas systems are often described as programmable molecular search engines. Give them a guide RNA, and they will find matching DNA. That description is elegant — but incomplete.
Hidden inside most CRISPR systems is a tiny molecular checkpoint: a short DNA sequence called the Protospacer Adjacent Motif, or PAM. For Cas12a, this PAM is typically TTTV (where V can be A, C, or G). If those four letters are not sitting directly beside the target sequence, Cas12a simply does not engage.
At the molecular level, PAM recognition is not just a label. It allows Cas12a to locally destabilize double-stranded DNA and stabilize an R-loop structure, where guide RNA pairs with target DNA while displacing one strand. Without PAM, this structural checkpoint fails — even if the target sequence perfectly matches the guide RNA.
For genome editing, this limitation is manageable. Scientists can choose another nearby site. But diagnostics does not offer that luxury. Disease mutations occur wherever biology places them. Many clinically important mutations — especially single nucleotide variants (SNVs) — sit in PAM-poor genomic regions.
For years, this meant something uncomfortable: some of the most medically important mutations were effectively invisible to CRISPR diagnostics. The TRACER study begins from this exact real-world limitation.
JACS_Pam-independent-crispr-cas…
The Mystery of PAM: Why Does CRISPR Need This Tiny Sequence?
If CRISPR is programmable, why does it need PAM at all? This question appears simple but touches the evolutionary origin of CRISPR itself.
CRISPR systems evolved in bacteria as immune defense systems against viruses (bacteriophages). When viruses infect bacteria, they inject foreign DNA. CRISPR systems store small pieces of this viral DNA inside bacterial genomes as “memory sequences.”
But this creates a dangerous problem. If CRISPR could cut DNA purely based on sequence matching, it might attack its own stored CRISPR memory sequences — essentially causing self-destruction.
PAM solves this.
In bacterial genomes: Stored CRISPR sequences do not contain PAM.
In viral genomes: Target DNA usually contains PAM.
So PAM acts like a self vs non-self identity tag.
Only DNA with PAM gets targeted.
This elegant evolutionary safety system became a limitation when CRISPR was repurposed for diagnostics and genome engineering.
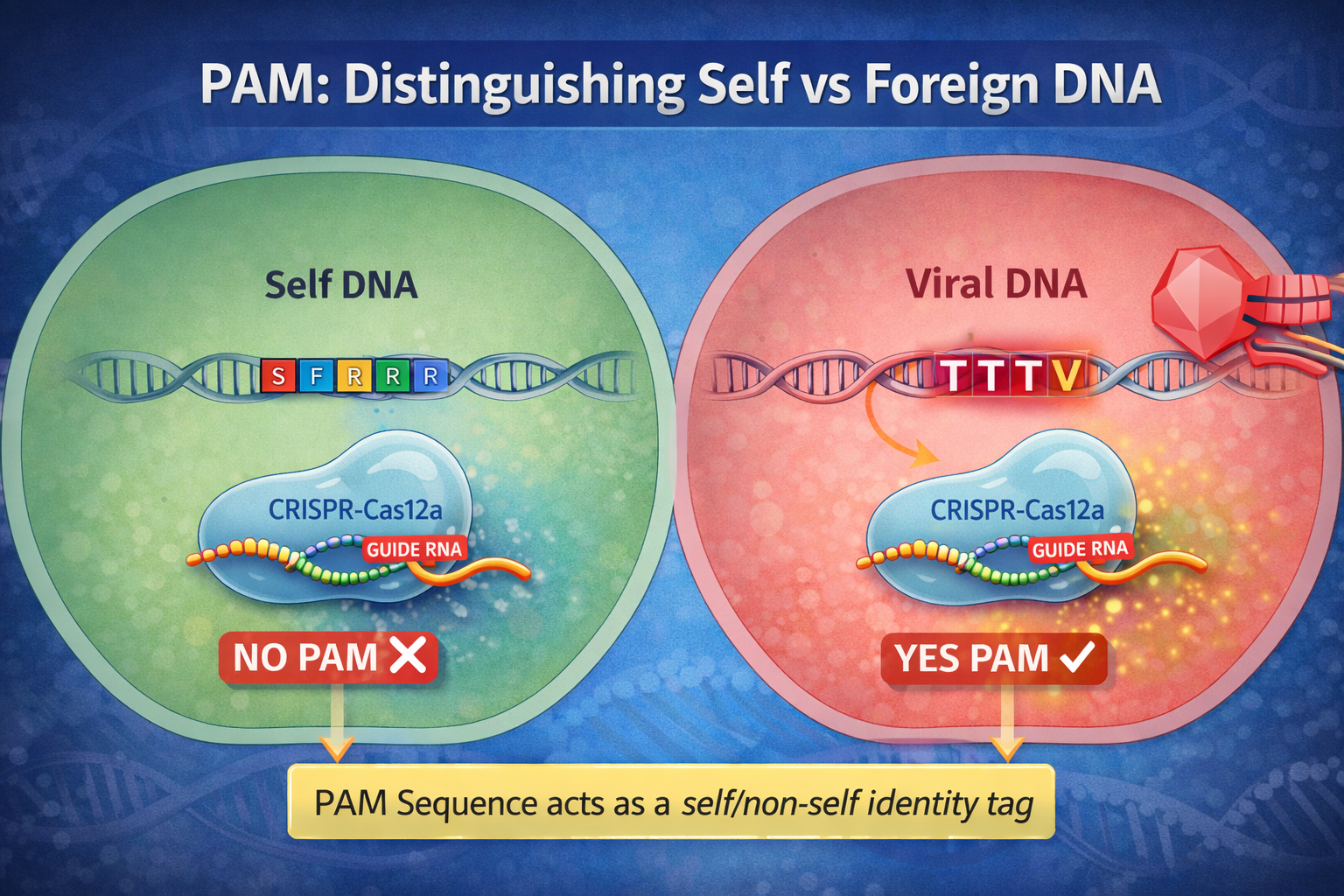
The Structural Reason PAM is Needed
Beyond evolution, PAM plays a physical role.
Cas12a cannot randomly open double-stranded DNA. DNA is tightly packed and energetically stable. PAM recognition gives Cas12a a foothold to begin unwinding DNA and testing sequence complementarity.
Without PAM: Cas12a cannot efficiently open dsDNA → No R-loop → No cleavage → No signal.
But here is the crucial detail: Cas12a does not need PAM when DNA is already in single stranded state.
This overlooked biochemical property is the foundation of TRACER.
Why CRISPR Became So Important in Diagnostics
Traditional mutation detection technologies each come with trade-offs. qPCR struggles with rare mutation detection in high wild-type background. Sequencing methods are accurate but expensive and slow.
CRISPR introduced something new: molecular recognition combined with signal amplification.
Cas12a has two cleavage modes. The first is cis-cleavage, where Cas12a cuts target DNA. The second is trans-cleavage, where activated Cas12a nonspecifically cuts nearby single-stranded DNA reporters, generating amplified fluorescence signals.
This is why CRISPR diagnostics can detect extremely low DNA concentrations. But the cascade depends on successful initial recognition — which normally requires PAM.
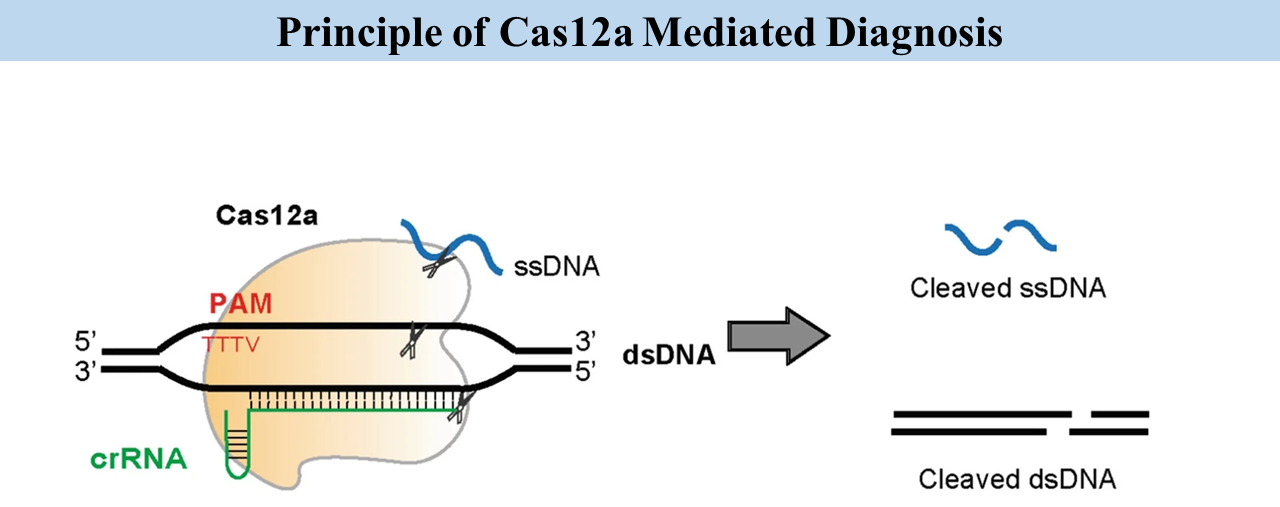
The Quiet Observation That Changed Everything
The TRACER story begins not with a new enzyme, but with a careful observation.
Cas12a is strict about PAM when interacting with double-stranded DNA. But when Cas12a encounters single-stranded DNA, PAM is no longer required.
This reframes the entire challenge. Instead of engineering new Cas proteins or inserting artificial PAM sequences, the authors asked a simpler question:
What if we don’t change Cas12a?
What if we change the DNA?
If double-stranded structure creates PAM dependency, converting DNA into single-stranded form might remove the restriction entirely.
That idea became TRACER.
TRACER: Changing the Rules Without Changing the Enzyme
TRACER (Mutant Target-Recognized PAM-Independent CRISPR-Cas12a Enzyme Reporting System) works by converting amplified DNA into a form Cas12a can recognize without PAM.
First, the target region is amplified using PCR or RPA. At this stage, DNA is still double stranded.
Next, the amplicons are converted into single-stranded DNA.
In BA-TRACER, alkaline treatment using NaOH denatures DNA strands, followed by neutralization using HCl.
In HC-TRACER, DNA is heat-denatured and rapidly cooled, preventing reannealing.
Both approaches generate abundant single-stranded DNA while preserving crRNA binding efficiency, ensuring strong Cas12a trans-cleavage activation.
Once DNA is single stranded, Cas12a becomes effectively PAM-independent.
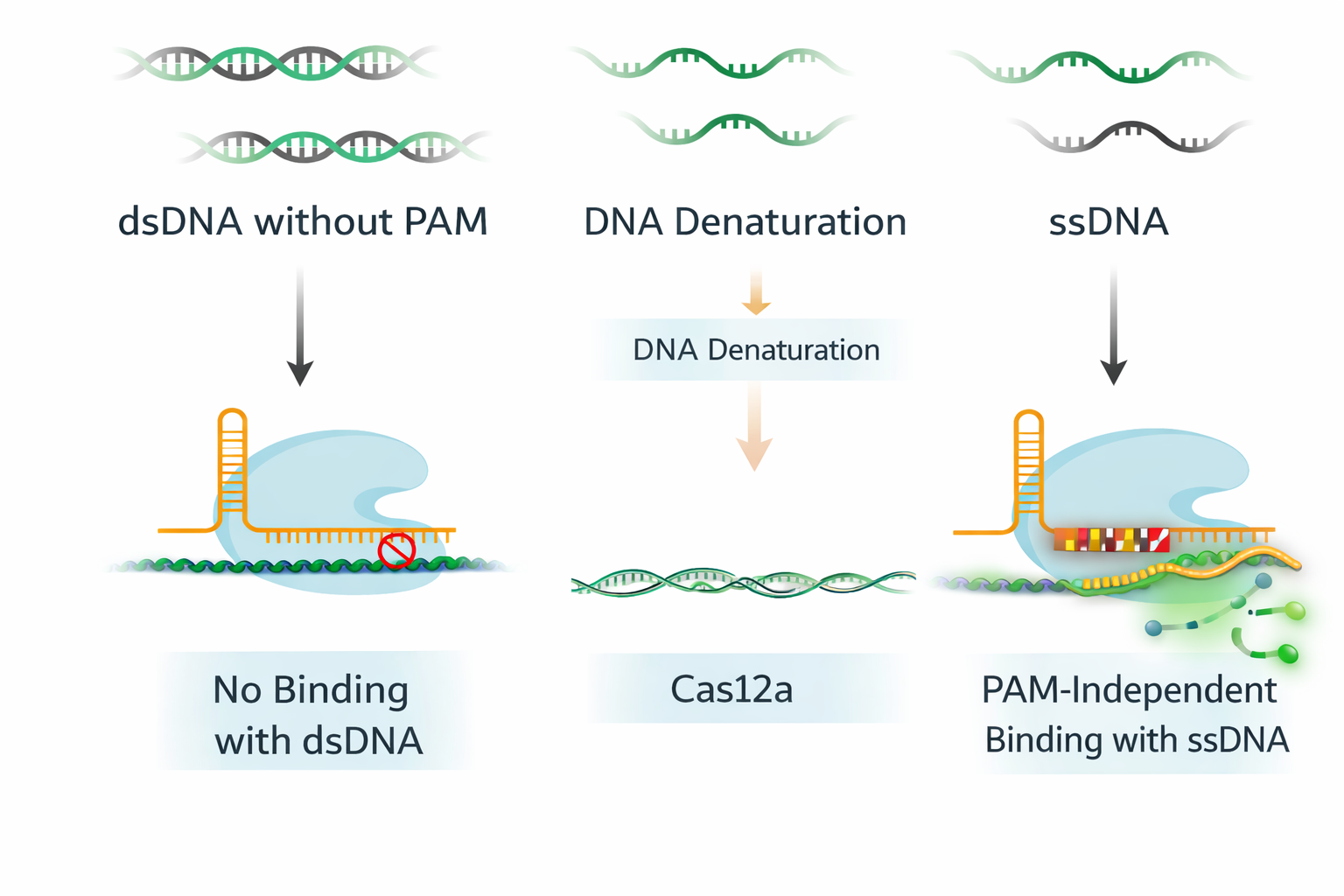
Experimental Proof: Demonstrating PAM-Independent Activation
PAGE and fluorescence assays confirmed that treated DNA behaves like single-stranded substrate. PAM-free DNA targets showed no Cas12a activity in native form but produced strong fluorescence after TRACER treatment.
This confirmed that PAM independence was achieved by changing DNA structure rather than modifying Cas12a itself.
Sensitivity: Detecting DNA at the Attomolar Scale
TRACER achieved detection limits near 0.5 attomolar, meaning extremely low DNA molecule counts can be detected.
PCR-based TRACER produced stronger signals than RPA, likely due to amplification efficiency differences.
This sensitivity is clinically meaningful for early mutation detection.
The Real Test: Detecting Single Nucleotide Variants
SNV detection depends heavily on the seed region of crRNA-target hybridization.
Mismatch penalties in the seed region reduce binding energy below the threshold required for Cas12a activation. The authors exploited this by designing crRNAs with strategic mismatch placement.
Using this strategy, TRACER detected mutant DNA even when it represented only 0.1% of total DNA population. This level of sensitivity is particularly important for liquid biopsy applications, where tumor-derived DNA fragments can exist at extremely low abundance within circulating cell-free DNA.
Clinical Validation: Performance in Real Tumor Samples
TRACER was tested on breast cancer (ESR1 mutation) and lung cancer (EGFR mutation) tumor samples.
For ESR1 mutation detection, TRACER achieved 100% sensitivity and specificity.
For EGFR mutation detection, accuracy exceeded 96%, with no false positives reported.
These results demonstrate strong clinical translation potential.
The Deeper Contribution: Substrate Engineering
Most CRISPR innovation focuses on protein engineering. TRACER represents substrate engineering — modifying the DNA environment rather than the enzyme itself.
This expands CRISPR targeting capability without requiring engineered enzyme variants. This approach reflects a broader trend in molecular diagnostics, where assay environment engineering is increasingly complementing protein engineering strategies.
Limitations of the Study
The workflow currently requires multiple steps, increasing contamination risk. Detection depends on upstream amplification efficiency. Clinical validation sample size remains moderate. crRNA design optimization is still required per mutation.
Future work will likely focus on one-pot PAM-independent CRISPR assays.
The Bigger Future: CRISPR Without Genomic “No-Go Zones”
Removing PAM dependency moves CRISPR diagnostics toward genome-wide mutation detection capability.
In liquid biopsy settings, where tumor DNA may represent extremely small fractions of circulating DNA, PAM-independent CRISPR detection could dramatically expand detectable mutation space.
Research Paper Decoded Insight
TRACER moves CRISPR diagnostics closer to what precision medicine truly needs — the ability to detect any mutation, anywhere, at extremely low abundance. By removing PAM as a targeting constraint, this approach expands the fraction of the genome that becomes diagnostically accessible. For cancers driven by rare mutations or monitored through liquid biopsy, this could directly influence how early relapse is detected and how quickly therapy decisions are made. If integrated into simplified clinical workflows, PAM-independent CRISPR could shift mutation detection from specialized labs toward routine diagnostic settings. In the long term, the real impact of TRACER may not be making CRISPR better — but making previously undetectable disease biology finally visible.
