Why You Need to Understand PCR
Whether you’re in your undergraduate molecular biology course, pursuing a master’s in biotechnology, or starting your PhD research, PCR (Polymerase Chain Reaction) is a non-negotiable foundational knowledge. You’ll encounter it in every lab rotation, every journal article on molecular biology, and likely in your own research projects.
But understanding PCR goes beyond memorizing the three steps. This article teaches you the why behind each step and practical decision-making that transforms PCR from a “recipe you follow” into a tool you can troubleshoot, optimize, and adapt to your research needs.
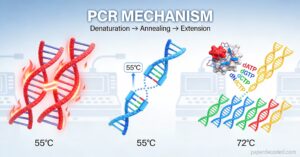

The Building Blocks – What PCR Actually Needs
As we all know PCR mimics DNA replication in a simplified, enzyme-minimal laboratory setup. In biological system e.g. our cells, replication is done with the involvement of multiple enzymes like DNA polymerase, DNA gyrase, DNA ligase, topoisomerase etc. But in laboratory setup inclusion of multiple enzymes for specific needs of replication is not a valid approach. Only the main enzyme DNA polymerase is used. Other needs are met by the thermal cycler or simply the PCR machine along with other raw material essential for polymerization.
Template DNA
The DNA containing your target sequence (genomic DNA, plasmid DNA, cDNA). Purity matters because common carryovers—phenol, EDTA, proteinase K, salts—directly inhibit amplification (check DNA purity with nanodrop).
Decoded Insight: If PCR fails, dilute the template. Recovery after dilution strongly suggested for removal of PCR inhibitors.
Primers (Specificity is Designed)
Primers (typically 18–25 bases) define where amplification starts and ends. Two primers are required:
- Forward primer: binds the antisense strand
- Reverse primer: binds the sense strand
Make sure to prepare 100 μM or 10 μM stock primer concentration.
Design principles that decide success:
- Tm: ~55–65 °C (use NEB Tm calculator based on the polymerase enzyme)
- GC: ~40–60%
- Avoid self-complementarity and primer–dimer formation (check with IDT oligo analyzer)
Decoded Insight: Most PCR failures trace back to primer design or annealing temperature—not the enzyme.
DNA Polymerase (The Thermostable Engine)
Standard Taq polymerase or other thermostable DNA polymerases like Q5, Phusion, withstands repeated heating and synthesizes DNA optimally near 72 °C.
When Taq is ideal: diagnostics, screening, routine amplification. When it isn’t: cloning or applications requiring high fidelity.
dNTPs: The DNA Building Blocks
dNTPs (deoxynucleotide triphosphates) are deoxy-versions of RNA building blocks, with three phosphate groups attached. Taq strips off two phosphates (releasing pyrophosphate) and incorporates the remaining nucleotide into the growing DNA strand. dNTPs are the main monomer which slowly consumed and polymerized to generate new DNA the polymeric form as PCR product.
Important for troubleshooting: Old dNTPs (>1–2 years at room temperature) degrade. If a PCR that always worked suddenly fails, suspect old dNTPs. Buy fresh ones or prepare fresh aliquot.
Buffer and Magnesium: The Chemical Environment
Taq works in a buffer solution containing Mg²⁺ ions. Think Mg²⁺ as the “glue” holding everything together:
Three critical roles of Mg²⁺:
Cofactor for Taq: Mg²⁺ binds to dNTPs, stabilizing them for incorporation
Stabilizes base pairing: Mg²⁺ shields negative charges on DNA backbone, allowing strands to stay together
Maintains primer-template stability: Low Mg²⁺ = primers fall off; high Mg²⁺ = primers stick too hard
Mg²⁺ optimization (crucial troubleshooting step):
Default: 1.5 mM MgCl₂
Too low (0.5–1.0 mM): Weak product, nonspecific bands disappear (specificity improves)
Too high (2.5–3.0 mM): Strong product but lots of nonspecific bands, primer-dimers
Pro tip: When your PCR produces multiple bands, first try increasing annealing temperature, then try decreasing Mg²⁺. This solves 80% of specificity problems.
The Three-Step Cycle (How PCR Works)
PCR is elegant because it repeats the same three steps 25–40 times. Let’s discuss the magical three steps.
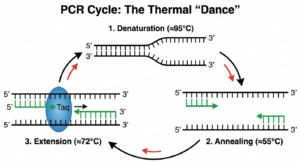
1) Denaturation (≈95 °C)
Heat separates double-stranded DNA into single strands by disrupting hydrogen bonds. This is necessary—without denaturation, primers can’t access the template.
Why it matters: Insufficient denaturation reduces available template; excessive time wastes enzyme activity.
Student mistake: “If 30 sec is good, 2 minutes must be better!” Wrong. Excessive heating denatures Taq (though it recovers in later cycles, but it hampers PCR efficiency). Stick to protocol.
2) Annealing (≈50–65 °C)
Primers bind their complementary sequences. This step determines specificity.
What physically happens:
Temperature is lowered, allowing primers to seek and bind their complementary sequences on the now-single-stranded template DNA. This is where specificity comes from.
The molecular dance:
Primers in solution rapidly associate/dissociate from template DNA (thousands of times per second)
When a primer finds its perfectly matching complement, hydrogen bonds form and it “sticks”
Mismatched primers (2-3 base) form weaker hydrogen bonds and dissociate during temperature transitions, preventing extension.
3) Extension (≈72 °C)
Taq polymerase extends primers 5′→3′, adding dNTPs to synthesize complementary strands.
Timing guide: ~1 minute per kilobase is sufficient for most targets.
Why Amplification Is Exponential
Each cycle doubles the number of target molecules (2ⁿ). Early cycles approach 100% efficiency; later cycles plateau as reagents deplete and competition increases.
Practical takeaway: 25–35 cycles are optimal. Beyond this, yield gains are minimal while artifacts increase.
Why extension time matters:
Too short: Product incomplete; you see faint band or no band
Too long: Wasted time; Taq isn’t slowing down, product is already complete
Way too long (10+ min): Taq degrades; faint bands appear
Troubleshooting Logic (Think, Don’t Guess)
No band: verify positive control → confirm template → check primers → run a gradient PCR → test for inhibitors.
Multiple bands: raise annealing temperature → reduce Mg²⁺ → redesign primers → use hot-start or touchdown PCR.
Faint band: increase template slightly → add a few cycles → refresh reagents.
Decoded Insight: Troubleshooting works best when you change one parameter at a time.
PCR Variants You’ll Encounter
- qPCR: real-time fluorescence for quantification
- RT-PCR: RNA templates via reverse transcription
- Multiplex PCR: multiple targets in one reaction
Transferable principle: All variants still depend on primer specificity, enzyme kinetics, and reaction chemistry learned here.
Research Paper Decoded Quick Take
PCR is simple in structure and deep in logic. Understand the chemistry behind denaturation, annealing, and extension—and PCR becomes predictable, fixable, and adaptable. This mechanistic foundation is the gateway to mastering Molecular Biology.
The best way to truly understand PCR? Run it 50 times. Make mistakes. Fix them. Understand why. That’s the real education. Maintain lab notebook.
Do you know there are different versions of PCR practiced in laboratories for specific experimental need and as a part of troubleshooting? Read This Decoded version of PCR Types.
Next in This Series
- PCR Primer Design Decoded
- PCR Optimization: Mg²⁺, Temperature, and Cycles
- PCR Troubleshooting Decoded
- PCR vs qPCR vs LAMP (Mechanisms Compared)

Pingback: PhD Student Position in Allergy and Asthma Research – Germany (LMU Klinikum) - Research Paper Decoded
Pingback: PAM-Independent CRISPR-Cas12a for Single Nucleotide Variant Detection || Paper Decoded - Research Paper Decoded