Research Paper Decoded breaks down high-impact research papers for UG, PG, and early PhD students—focusing on concepts, not jargon.
Research question
Can we use CRISPR-Cas12a to detect single-nucleotide variants (SNVs) in double-stranded DNA even when there is no suitable PAM sequence near the mutation, without modifying the enzyme itself?
Many clinically important SNVs are poorly positioned for standard Cas12a detection because they lack the canonical TTTV PAM nearby. This paper asks whether that limitation can be bypassed by changing the assay strategy rather than the protein.
Original Article Details:
Title: PAM-Independent CRISPR-Cas12a System for Specific Assays of Single Nucleotide Variants
Authors: Jinlong Ai, Jinhai Deng, Jingjing Hu, Xingxiang Pu, Tongyan Yuan, Yuling Teng, Han Li, Bojie Chen, Jinlian Du, Ling Jiang, Xiaoyan Chen, Erhu Xiong, and Ronghua Yang
DOI: https://doi.org/10.1021/jacsau.5c00011
Journal: JACS Au
Why should students care about this paper?
A single-nucleotide variant (SNV) is a change in just one DNA base, but that one change can:
- Make cancers resistant to therapy
- Influence prognosis
- Determine which drug a patient should receive
Detecting such mutations is especially hard when the mutant DNA is present at low levels among mostly normal DNA. This paper presents a promising strategy to use CRISPR for mutation detection without PAM sequence restrictions, representing an important conceptual advance in molecular diagnostics.
The basic problem with CRISPR-Cas12a diagnostics
Normally, Cas12a efficiently recognizes double-stranded DNA targets only when a short PAM sequence (typically TTTV) is present next to the target.
This creates three practical problems:
- Many clinically relevant SNVs do not have a suitable PAM nearby
- crRNA design becomes highly restricted
- Some important mutations become effectively invisible to standard CRISPR-Cas12a assays
In short, diagnostic success often depends more on genomic context rather than on clinical importance.
The simple idea behind TRACER
The authors introduce a strategy called TRACER.
Instead of modifying Cas12a, they modify the form of the DNA target.
Key insight:
LbCas12a can recognize single-stranded DNA (ssDNA) using sequence complementarity alone, without requiring a PAM.
By converting amplified double-stranded DNA into ssDNA just before detection, the strict PAM requirement can be bypassed while still using mutation-specific crRNAs.
Two ways to make single-stranded DNA
The paper describes two complementary approaches:
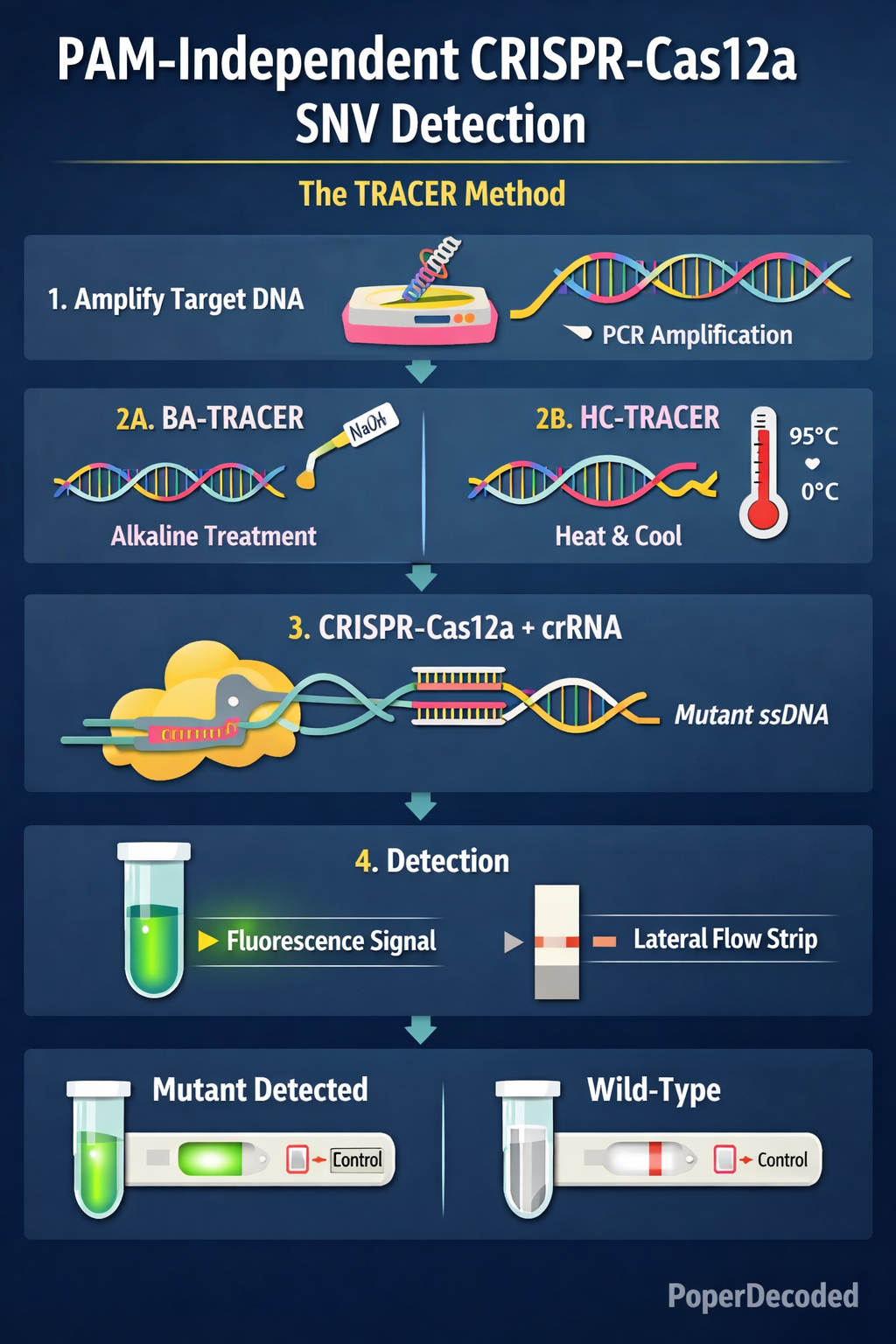
- BA-TRACER: Briefly treats amplicons with dilute NaOH to separate DNA strands, followed by neutralization so Cas12a remains active.
- HC-TRACER: Heats DNA to 95 °C to melt the strands, then rapidly cools to 0 °C to keep the DNA single-stranded long enough for detection.
Both methods generate ssDNA immediately before CRISPR detection, minimizing degradation.
Step-by-step workflow (student-friendly)
- Amplify the target DNA (PCR or RPA)
- Convert dsDNA into ssDNA (chemical or heat-based method)
- Add Cas12a and mutation-specific crRNA
- Cas12a is activated only if the mutant sequence is present
- Activated Cas12a cleaves a fluorescent reporter
After amplification, the CRISPR reaction is monitored for up to 90 minutes, with clear mutant–wild-type discrimination often visible by around 45 minutes.
Sensitivity and specificity
The assay shows strong analytical performance under controlled conditions:
- Sensitivity: Detection down to the attomolar range (0.5 aM with PCR)
- Allele discrimination: Can distinguish mutant DNA even when it represents ~10% of a mutant–wild-type mixture
- Specificity: Strong separation between mutant and wild-type signals
These results demonstrate high analytical sensitivity, although clinical performance still depends on sample type and workflow integration.
Does it work on real patient samples?
The authors tested the method on DNA extracted from FFPE tumor tissues:
- Breast cancer samples with ESR1 Y537S mutations
- Lung cancer samples with EGFR S768I mutations
In this small cohort, results showed high concordance with Sanger sequencing, with no false positives and a single missed EGFR mutant sample, indicating good but not perfect clinical sensitivity.
Limitations & future directions
1. Multi-step workflow
The assay involves several open-tube steps. Each step increases aerosol contamination risk, especially in high-throughput settings. Future designs should aim for closed-tube or one-pot formats.
2. Chemical and thermal handling
BA-TRACER uses simple NaOH/HCl steps that may be easier to implement than precise temperature control, while HC-TRACER requires rapid cooling. Choice of method may depend on laboratory infrastructure.
3. Timing of ssDNA generation
Single-stranded DNA can re-anneal quickly. In this assay, ssDNA is generated immediately before detection and kept cold (HC-TRACER) to reduce this issue, making timing rather than stability the key challenge.
4. crRNA optimization
Multiple crRNAs were tested for each target to achieve strong SNV discrimination, indicating that empirical optimization is still required.
5. Limited clinical scope
Only two cancer-related SNVs were evaluated using tumor tissue DNA. Broader validation across more mutations, diseases, and sample types (e.g., blood or plasma) is needed.
6. Point-of-care limitations
Best performance relies on PCR and fluorescence readout, which limits immediate use in true point-of-care settings.
Future work will likely focus on integrating amplification, strand separation, and CRISPR detection into a single, closed reaction to simplify the workflow and improve robustness.
Paper Decoded take-home message
This paper highlights a powerful design lesson:
When a protein’s native rules become limiting, changing the substrate or assay context can be just as effective as engineering the protein itself.
By converting dsDNA into ssDNA, the authors remove PAM constraints and expand what Cas12a-based diagnostics can detect. While TRACER is not yet a ready-to-use diagnostic kit, it provides a clear conceptual framework that can inspire future CRISPR assay design.
A question for you (pause before scrolling)
Imagine you are designing a CRISPR-based diagnostic for a real patient.
You know the mutation is important.
You know CRISPR can detect it.
But there is no PAM site nearby.
What would you try next?
- Change the enzyme?
- Change the crRNA design?
- Or change the form of DNA, as done here?
Try to justify your choice in 2–3 sentences. Your reasoning may help another reader see the problem differently.
Research Paper Decoded breaks complex research into concepts you can actually understand.
